Introduction
There are approximately 10,000 named species of microbes. It is estimated that there are between 10,000 and 100,000 more unidentified species for every identified one. Not only are there many types of bacteria, there are a lot of individual bacteria. A single spoonful of soil can have 100 million individual bacteria. A scraping of your gums can yield 1 million bacteria per cm2 (a cm2 is about the size of your little fingernail). The bacteria in and on our bodies makes up about 10% of our dry body weight.
Most of the currently known species of bacteria have been identified using traditional microbiological techniques such as the gram stain reaction, morphology, and metabolic reactions. Bacteria rarely live alone but in communities with other bacteria. This is true both in the environment and in and on our bodies. This class focuses on the role of bacteria in disease. Isolating a single bacterium species is the first step in identifying the bacteria possibly responsible for a disease process.
The first requirement for physically isolating a bacterium is that it can be cultured in the laboratory. This requires knowledge of optimal temperature for growth, optimal oxygen requirements, and optimal nutritional needs. We work with a very limited number of bacteria in this course. The bacteria we work with are also very easy to culture in the lab. Most bacteria are not this agreeable!
There are two main ways to isolate organisms.
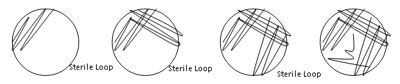
- Streaking for isolation on an agar plate
- The pour plate method
Streaking for isolation on an agar plate involves the successive dilution of organisms until you have the cells at a low enough density that single cells are physically isolated spatially to give rise to recognizable individual colonies. In the pour plate method, you dilute your sample sufficiently before you add it to molten cooled agar and then pour this mixture in a dish. The isolated cells give rise to individual colonies growing in the agar itself. This technique can be a little tricky. If the melted agar is too hot you kill all the bacteria. If the melted agar is too cool you end up with a big lump in your Petri dish. The streaking method yields individual colonies on the surface of the agar. This technique is much faster and easier to master.