Overview
You will be using a wide variety of media and biochemical tests to isolate and identify your unknown organisms. Your text has a good section on enrichment, selective, and differential media. Please review it before getting started.
You are only working with a small subset of the available media and tests. It is enough however to give you a good idea of the problem solving that goes on behind the scenes to support patient care.
The information provided on these pages was derived from the DIFCO Manual of media, which is also available in the lab.
Read the directions carefully.
You must pay close attention to the conditions of the tests and media to avoid erroneous results.
Things to watch for include:
- type of media you pull your sample from (enrichment or selective).
- mixed or pure cultures for inoculation.
- type of inoculation - one or more colonies, needle prick from center or whole colony, light or heavy innoculation.
- application - do you stab, streak or smear the plate or tube.
- length, temperature, and atmospheric conditions for incubation; some tests run for longer times if you get a negative result in 24 hours, others need to run for 4-5 days, others must be read at 24 hours because the colors change with storage.
Bacitracin/SXT Sensitivity
Intended Use
Bacitracin differential disks are used to presumptively identify Group A, beta-hemolytic streptococci from other beta-hemolytic streptococci. The combination of SXT sensitivity increases the accuracy of the results.
Principle
Bacitracin is an antibiotic isolated from Bacillus subtilis. It inhibits cell wall synthesis mainly through inhibiting the biosynthesis of peptidoglycan. SXT inhibits folate metabolism which interferes with bacterial DNA synthesis. Group A, beta-hemolytic streptococci are more sensitive to bacitracin than other beta-hemolytic streptococci.
Test Procedure
The standard protocol has been modified for our lab.
- Using a loop, select 3-4 well isolated colonies, ideally from an 18-24 hour culture. Transfer to a small amount of sterile water.
- Adjust the turbidity to 0.5 McFarland standard.
- Use the procedure outlined in antimicrobial susceptibility testing to swab the entire plate to obtain confluent growth.
- Visually divide the plate in thirds, place a bacitracin and SXT in their section of the plate. Using sterile forceps or a swab, lightly but firmly press the disks to the agar surface to adhere them.
- Save the other section for the optochin disk.
- Invert the plates and incubate them for 18-24 hours at 35°C in 5-10% CO2.
- Incubate another 24 hours if the results are negative.
Results
- Any zone of inhibition around the disk is considered sensitive (S).
- No zone of inhibition with growth up to the disk is considered resistance (R).
This table is from MacFaddin, Biochemical Tests for Identification of Medical Bacteria.
| Bacitracin | SXT | Presumptive ID |
|---|---|---|
| S | R | Group A b-streptococci |
| R | R | Group B b-streptococci |
| R | S | Not Group A or B b-streptococci |
| S | S | Rule out Group A or B with serologic tests |
Limitations
- Only beta-hemolytic streptococci should be tested.
- While this test is accurate it is not highly specific. Other biochemical or serological tests are required for accurate identification.
- The growth should be confluent. Too light of a growth could cause some non-group A streptococci to appear susceptible to bacitracin.
Catalase
Intended Use
- Differentiates Streptococcus (-) from Micrococcus (+)
- Differentiates Staphylococcus (V+) and Bacillus (+) from Clostridium (-)
Principle
Hydrogen peroxide (H2O2) is the end product of aerobic breakdown of sugars. Since it is toxic to bacterial cells, most aerobic bacteria produce catalase or peroxidase to protect themselves. Streptococcus, Enterococcus, and Lactobacillis are exceptions. Since they do not use the cytochrome c pathway, they do not produce H2O2 and lack catalase.
Test Procedure
- Transfer a well isolated colony to a clean glass slide and add 1 drop of 3% H2O2.
- Do not reverse the order and do not mix.
- Observe for immediate bubble formation.
- Use 15% H2O2 for the detection of catalase in anaerobes.
Results
- The formation of bubbles is considered a positive result.
Limitations
- Do not take your colony from a blood agar plate. The catalase present in the erythrocytes will give a false positive result.
- H2O2 is unstable. You can do a quality control test of the H2O2 reagent by placing a drop on a blood agar plate. Vigorous bubbling should result.
Coagulase Test
Intended Use
Differentiates Staphylococcus aureus from other Staphylococcus species.
Principle
The coagulase test detects the presence of free and bound staphylcoagulase. This enzyme is excreted extracellularly by human strains of Staph. aureus. The mechanism of action is unknown.
Test Procedure
- Thaw a tube of 0.5 mL rabbit plasma.
- Inoculate a loop-full of organism into the tube. Chose a well isolated colony.
- Ideally you should incubate the tube at 35°C for 4 hours checking every 30 minutes for clot formation. We incubate them overnight and put them in the refrigerator until the next lab period with comparable results.
- Check for clot formation.
- Dispose of the tube in the biohazard container.
Results
The formation of a clot in the bottom of the tube is considered a positive result. The clot will not move as you tilt the tube. Unclotted plasma will flow in the tube.
Limitations
- Methicillin resistant Staph. aureus have reduced clumping factor.
- Do not shake or agitate the tube as this could break up the clot.
- Some staphylococci strains produce fibrolysin after prolonged incubation at 35°C that can break up the clot resulting in false negative. Incubate the tube overnight at room temperature if you do not get a clot in 4 hours.
- Some other rarely encountered staph species are also coagulase positive by the tube method.
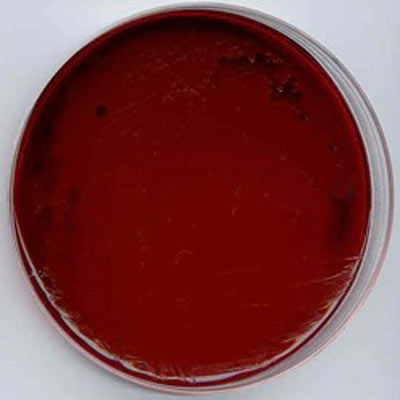
Eosin Methylene Blue (EMB) Agar
Eosin Methylene Blue (EMB) Agar
Principle
A differential plating medium for the detection & isolation of the gram-negative enteric bacteria.
Purpose
- To aid in the differentiation of lactose fermenting bacteria from non-lactose fermenting bacteria.
- To aid in the differentiation of Enterobacter aerogenes and Escherichia coli.
Test Procedure
- Inoculate the organism directly onto the surface of an EMB agar plate and streak for isolation.
- Incubate inoculated plate aerobically at 37°C.
- Examine for growth after 18-24 hours of incubation.
Interpretations
- Coliforms that utilize the lactose and/or sucrose are blue/black with a greenish metallic sheen. Indicative of Escherichia coli.
- Coliforms such as Klebsiella pneumoniae have mucoid colonies that may be purple and/or exhibit a green metallic sheen.
- Good to excellent, colorless colonies indicative of Proteus vulgaris, Salmonella choleraesuiss, and Shigella spp.
Fluid Thioglycolate Medium (FTM)
Tests the oxygen requirements of different microorganisms.
Intended Use
Characterizes microbes according to their oxygen requirements
- Obligate aerobes
- Facultative anaerobes
- Obligate anaerobes
Principle
Various types of bacteria require various oxygen (or oxygen-free) environments to grow in.
Test Procedure
- Dilute your organism in a tube of sterile water to obtain a turbidity equivalent to the 0.5 McFarland test standard. Hold your diluted tube and the 0.5 McFarland test standard against the black-lined McFarland reference card to accurately rate the turbidity.
- Using a sterile 1mL pipette, place 1 mL of organism into the middle of the tube.
- Cap tightly; do not jostle.
- Incubate for 24 hours at 37°C.
Results
- Strict (obligate) aerobes grow at the surface of the medium where there is a high concentration of oxygen.
- Obligate anaerobes grow near the bottom of the broth tube where there is no oxygen.
- Facultative anaerobes grow best where more oxygen is present, but growth will occur throughout the broth tube.
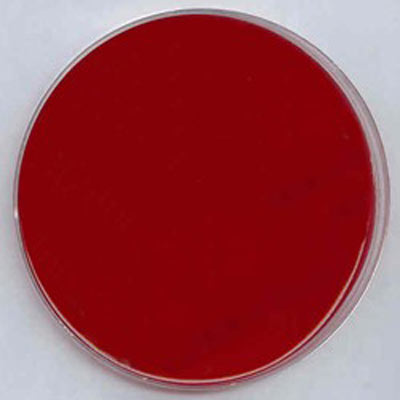
Hemolysis (Blood) Agar
Hemolysis - Blood Agar
Intended Use
Blood agar is used to support the growth of fastidious organisms and to determine the type of hemolysis (destruction of red blood cell walls) an organism produces.
Principle
Blood agar is a rich medium that has been supplemented with fresh 5-10% blood. The hemolytic response can be dependent upon the type of blood. Sheep blood is commonly used, but some organisms require rabbit or bovine blood.
Test Procedure
- Streak a plate of blood agar for isolation.
- Optional: Do your last streak with a needle and poke into the agar. This usually gives clear, reliable zones of beta hemolysis and is especially important to see the effects of streptolysin O which is oxygen labile. See page 84 of the Difco/BBL Manual.
- Incubate the plates at 37°C for 24-48 hours. Strep organisms should be incubated in the CO2 incubator.
- The plate will be a brownish red color after 48hours.
Results
You can differentiate four types of hemolysis by the appearance of the agar.
- Beta hemolysis is indicated by a clear colorless zone surrounding the colonies. There has been total lysis of the red blood cells.
- Alpha hemolysis is indicated by a small zone of greenish to brownish discoloration of the media. This is caused by the reduction of hemoglobin to methemoglobin and its subsequent diffusion into the surrounding medium.
- Alpha prime hemolysis is indicated by a zone of complete hemolysis, surrounded by a zone of partial hemolysis, a pink halo. This pattern can be easier to see if you scrape off the colony.
- Gamma hemolysis is indicated by no change in the media.
Limitations
- The patterns of hemolysis can vary with the incubation atmosphere and the type of blood in the media.
- Some Staph organisms will only show hemolysis after they have been refrigerated following incubation.

Hippurate Hydrolysis
Hippurate Hydrolysis
Intended Use
Aids in the differentiation of β-hemolytic Strep. agalactiae from other β-hemolytic streps. It should not be used with all streps, only the β-hemolytic streps of human and bovine origin. It is critical for the identification of Camplyobacter jejuni.
Principle
Hippurate acid is hydrolyzed to benzoic acid and glycine by the enzymatic action of hippuricase. The glycine end product is detected by the addition of ninhydrin reagent.
Test Procedure
- Add 4-6 drops of sterile water to a small microcentrifuge tube.
- Heavily inoculate the microcentrifuge tube.
- The tube should be cloudy looking after inoculation, turbid.
- Add a Hippurate disk to the tube with sterile forceps.
- Incubate in a 37°C for 24 hours.
- Carefully add 2-3 drops of the ninhydrin solution down the side of the tube to form an overlay over the cell mixture.
- Do NOT mix the solutions.
- Set aside for 5-10 minutes.
Results
Development of a deep purple color within 5-10 minutes is a positive result.
Limitations
- A faint purple color is considered a negative result.
- This test is only useful when distinguishing β-hemolytic streptococcus of human or bovine origin.
- The hippurate solution deteriorates in 7 days at 4°C.
- The ninhydrin solution deteriorates in 6 months.
- Hippurate hydrolysis is also found in some species of other genera, such as Bacillus, Corneybacterium, Enterobacteriaceae, and others.
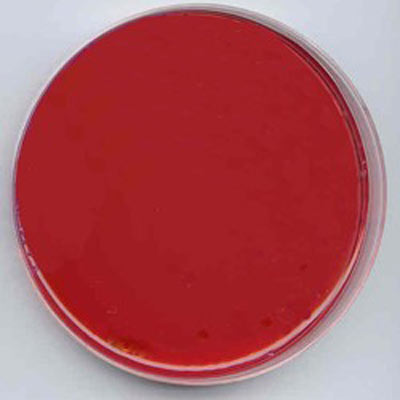
MacConkey (Mac) Agar
MacConkey (Mac) Agar
Principle
- A differential plating medium recommended for use in the isolation and differentiation of lactose-fermenting organisms from lactose non-fermenting gram negative enteric bacteria.
- Selectivity of the medium is due to the presence of crystal violet and bile salts which markedly to completely inhibit the growth of gram positive organisms.
Purpose
- Organisms capable of fermenting lactose produce a localized pH drop which, followed by the absorption of neutral red, imparts a red/pink/purple color to the colony. A zone of precipitated bile may also be present due to this localized drop in pH.
- Organisms that do not ferment lactose remain colorless and translucent. They are easily detected by transmitted light and appear as colorless colonies against a red background.
Test Procedure
- Inoculate the organism directly onto the surface of a MacConkey agar plate and streak for isolation.
- Incubate inoculated plate aerobically at 35-37°C.
- Examine for growth after 18-24 hours of incubation.
Interpretation
- Good to excellent growth, red/pink/purple colonies with bile precipitate indicative of Escherichia coli.
- Good to excellent growth, red/pink/purple colonies without bile precipitate indicative of Enterobacter aerogenes.
- Good to excellent, colorless colonies without bile precipitate indicative of Proteus vulgaris, Salmonella typhimurium, and Shigella spp.
Motility Test Medium
Intended Use
- Used for detecting motility of microorganisms.
Principle
Motility is apparent by the presence of diffuse growth away from the line of inoculation.
Test Procedure
- Inoculate with growth from an 18-24 hour culture by stab inoculation with a needle.
- Incubate at a temperature and duration appropriate for the organism being tested.
- Examine tubes for growth and signs of motility.
Results
- Motility is apparent by the presence of diffuse growth away from the line of inoculation.
- Non-motile organisms only grow along the line of inoculation.
Limitations
- The motility of Proteus spp. is temperature dependent.
- Due to the temperature dependency of motility in some organisms, a negative tube should be incubated an additional 5 days at a lower temperature of 22-25°C.
Mannitol Salt Agar (MSA)
Principle
Selective and differential media used for the isolation of Staphylococcus aureus.
Intended Use
Differentiates Staphylococcus aureus (+) from other Staphylococcus species.
Principle
- Contains 7.5 % NaCl (salt) which inhibits the growth of most microorganisms, except Staphylococcus species.
- Contains phenol red indicator which is red/pink under neutral/basic conditions, but turns yellow under acidic conditions.
- When mannitol is fermented there is a decrease in pH, turning the red/pink media yellow.
Test Procedure
- Inoculate the organism directly onto the surface of an MSA plate and streak for isolation.
- Incubate inoculated plate aerobically at 37°C.
- Examine for growth after 18-24 hours of incubation.
Results
The yellowing of the red/pink media indicates a positive result.
Novobiocin
Intended Use
Novobiocin Differentiation Disks are useful for presumptively distinguishing Staphylococcus saprophyticus from other coagulase-negative staphylococci (CoNS) in clinical specimens.
Principle
Novobiocin Differentiation Disks are prepared by impregnating 5µg of novobiocin onto high quality 6mm diameter filter paper disks. Novobiocin Differentiation Disks are useful in presumptively distinguishing S. saprophyticus from other CoNS. Novobiocin inhibits the synthesis of DNA and RNA. A zone of inhibition is produced by contact with the novobiocin.
Procedure
- Allow disks to equilibrate to room temperature.
- Using a pure 18-24 hour culture, prepare a suspension, equivalent to a McFarland 0.5 opacity standard, in Tryptic Soy Broth, Sterile Water, or Brain Heart Infusion (BHI) Broth.
- Inoculate Mueller Hinton Agar, Blood Agar, or Tryptic Soy Agar (TSA) plate with a sterile swab to obtain confluent growth.
- Aseptically apply one novobiocin disk onto the inoculated agar surface and lightly press down to ensure full contact with the medium.
- Incubate aerobically for 18-24 hours at 35-37 degrees C.
- Measure (in millimeters) the diameter of the zone of inhibition around the novobiocin disk, and record as susceptible or resistant.
Interpretation of Results
- Sensitive - A zone of inhibition greater than 16mm
- Resistant - A zone of inhibition less than or equal to 16mm
Limitations
- It is recommended that biochemical and/or serological tests be performed on colonies from pure culture for complete identification.
- The novobiocin disk is not helpful and can give misleading results if it is performed on isolates other that those from urinary specimens.
- Occasional human isolates that are not S. saprophyticus, S. cohnii subsp., or S. xylosis may also be resistant to novobiocin.
Oxidase Biochemical Assay
Tests for the presence of the enzyme indophenol oxidase.
Intended Use
The oxidase test is based on the production of an enzyme called indophenols oxidase. This enzyme oxidizes a redox dye (present in the reagent) which results in a color change of yellow to dark purple.
Principle
Indophenol oxidase, in the presence of atmospheric oxygen, oxidizes the phenylenediamine oxidase reagent to form a dark purple compound, indophenol.
Test Procedure
- Have your instructor or IA crush the ampule inside the dropper.
- Tap bottom on tabletop a few times. Then invert for convenient drop-by-drop dispensing of reagent
- Preparation for testing:
- Colonies to be tested must be isolated from other colonies
- The use of fresh isolates (18-24 hr cultures) is recommended for routine testing.
- If refrigerated, cultures must be allowed to reach room temperature prior to testing
- Performing the test – Filter Paper Method
- Add a few drops of oxidase test reagent to a strip of filter paper (Whatman No. 1 or equivalent).
- Streak a loopful of bacteria onto the reagent-saturated paper with a platinum loop or wooden applicator stick. Use of steel of nichrome loops may cause false-positive reactions
Results
Positive reactions turn the bacteria violet to purple immediately, or up to 30 seconds. Negative reactions remain colorless or turn light pink/light purple after 30 seconds. Delayed reactions should be ignored.
Limitations
- Allow up to 30 seconds for a positive reaction.
- Any delayed reactions should be considered negative.
- Do not add excess reagent, at it may cause the reaction to fade on oxides-positive organisms.
- Steel loop, nichrome loop, and wire loop containing iron may give a false-positive reaction. A platinum loop or wooden applicator stick is recommended.

Salt Tolerance Broth
Salt Tolerance Broth
Intended Use
Salt tolerance broth is intended to differentiate non-beta-hemolytic strains of streptococci.
Principle of Use
Brain Heart Infusion (BHI) broth is supplemented with 6.5% sodium chloride and bromcresol purple as a pH indicator. The indicator is included to make reading the test results easier. The broth also includes dextrose. The fermentation of dextrose (glucose) results in the production of acid. This changes the pH of the media causing the media to turn from purple to yellow.
Test Procedure
- Select no more than 2-3 colonies (preferably from an overnight culture) to inoculate a tube of salt tolerance broth.
- It is important to lightly inoculate the tube otherwise you may get a false positive.
- Loosen the cap and incubate aerobically for 24 hours at 37°C.
- Continue incubation up to 72 hours if you get a negative result at 24 hours.
Results
A positive reaction is indicated by obvious turbidity in the media with or without a color change. A negative result is indicated by no growth after 72 hours. Enterococcus spp. typically changes the media color within 24 hours.
Limitations
- Many staphylococci can grow in media containing 10% salt. Mannitol salt agar has 7.5% salt.
- Salt tolerance media was intended to differentiate catalase negative gram-positive cocci. Be sure to perform a catalase test before you proceed with the salt tolerance broth test.
- Other species of catalase negative gram-positive organisms can grow in this media.
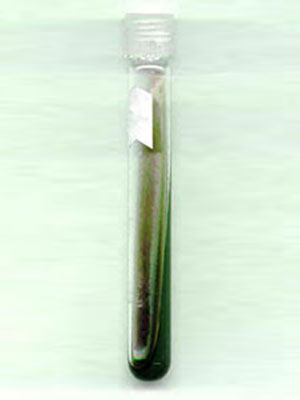
Simmons Citrate Agar Slant
Simmons Citrate Agar Slant
Principle
Used for the differentiation and identification of Enterobacteriaceae on the basis of citrate utilization, citrate being the sole carbon source.
Purpose
Colonies capable of utilizing citrate as a carbon source produce a local increase in pH, changing the color of the medium from green to blue. Only citrate positive organisms will grow on this medium.
Test Procedure
- Inoculate the organism directly onto the surface of a Citrate slant.
- Incubate aerobically at 35-37°C.
- Examine for growth and color change after 18-24 hours of incubation.
Interpretations
Good growth with the medium color turning blue indicative of Enterobacter aerogenes and Salmonella choleraesuis.






